
LARGE SCALE DISCRIMINATION BETWEEN NEURAL MODELS AND EXPERIMENTAL RECORDINGS
Professor Rick Gerkin1, Professor Sharon Crook2, Russell Jarvis3
1 Department of Mathematics and Statistics, Arizona State University, USA
2 Department of Mathematics and Statistics, Arizona State University, USA
3 Department of Life Sciences, Arizona State University, USA

nocite: | @[Brette, Romain and Gerstner, Wulfram], @[Pack, Christopher C. and Berezovskii, Vladimir K. and Born, Richard T.] ---
Abstract
NeuroML-DB [2] catalogues over 1,500 published models obtained in NeuroML format from Open Source Brain [5]. Complementing OSB, NeuroML-DB provides systematic characterizations of model complexity, electrophysiology, and morphology, making it easy to find, evaluate, and reuse models and their components
Example Features
We unified three different feature extraction protocols to create a very high dimensional feature space.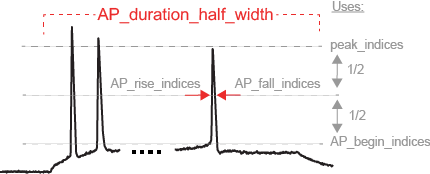
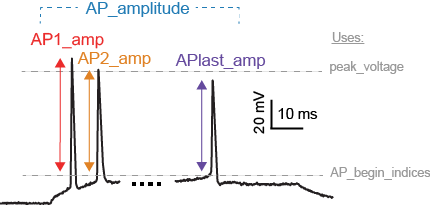
Modeling Methods
Publications Associated with Model Sources:
| Large Scale Model | Blue Brain Project (1035) | Gouwens et al (2018) [6] |
|---|---|---|
| Publication | Markram et al (2015) [8] | Gouwens et al (2018) [6] |
Extracted Feature Sources:
| Feature1 | Feature2 | Feature3 |
|---|---|---|
| Ephys Feature Extraction Library | allensdk.ephys.ephys_features | Druckmann et al. (2012) [3] |
Virtual Experiment Three Step Protocol Stimulate for 2s
| Injection 1 | Injection 2 | Injection 3 |
|---|---|---|
| at 1.0×Rheobase | at 1.5×Rheobase | at 3.0×Rheobase |
Results
Histogram to query Model/Data agreement
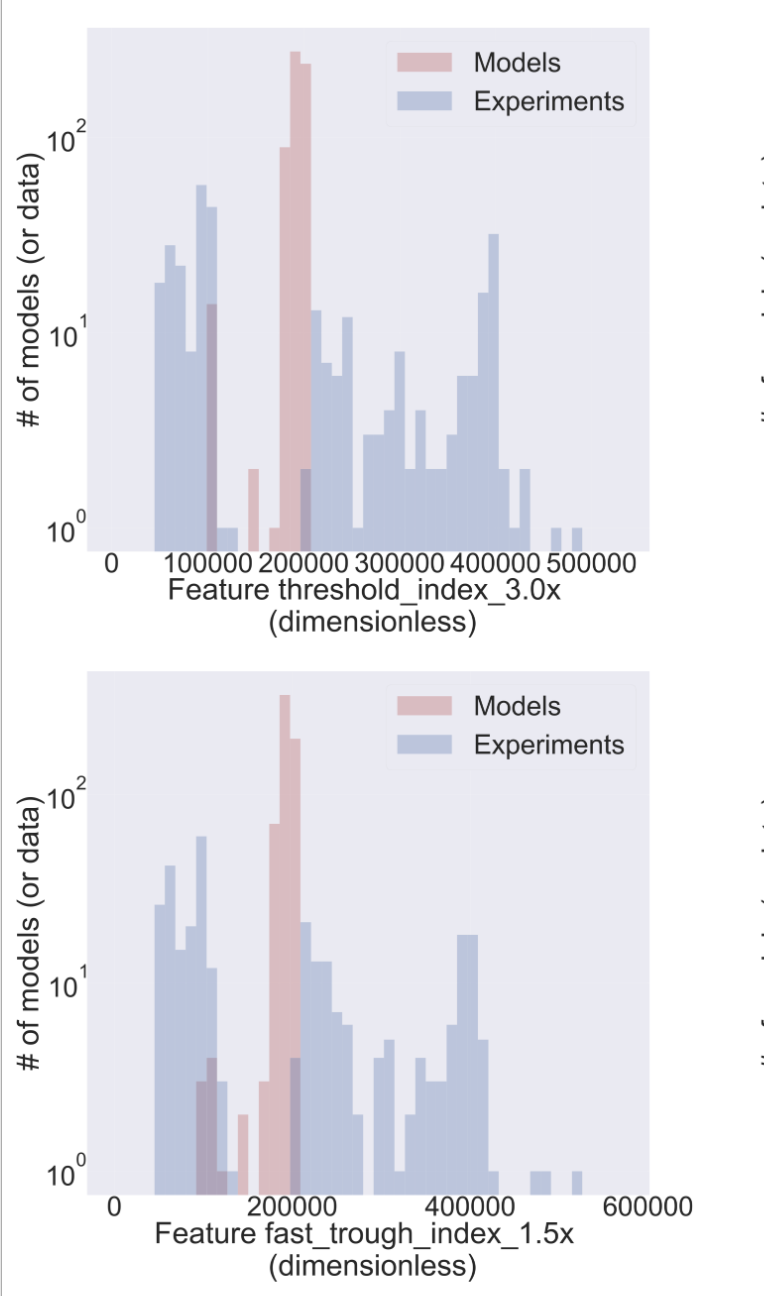
PCA Embedding
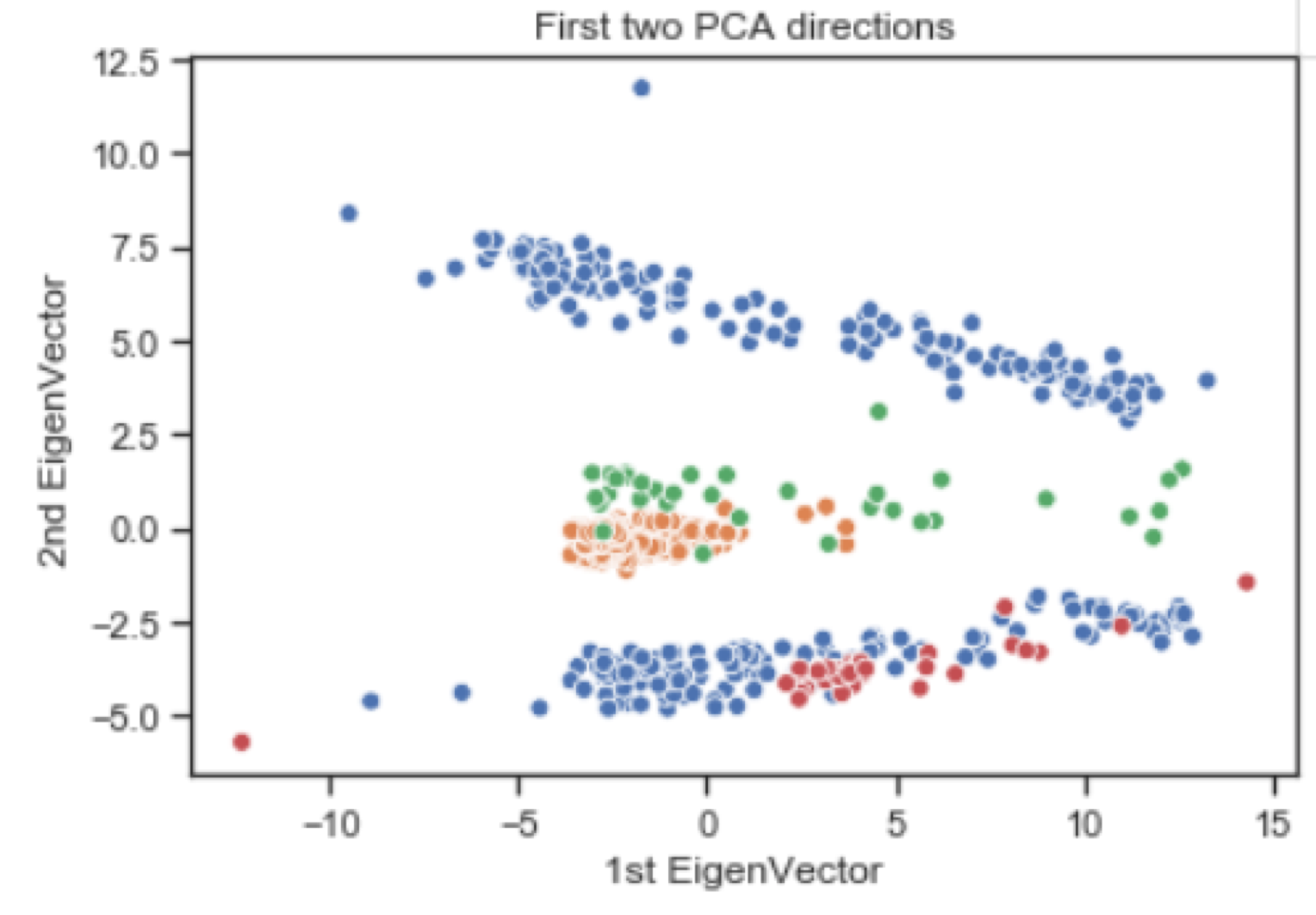
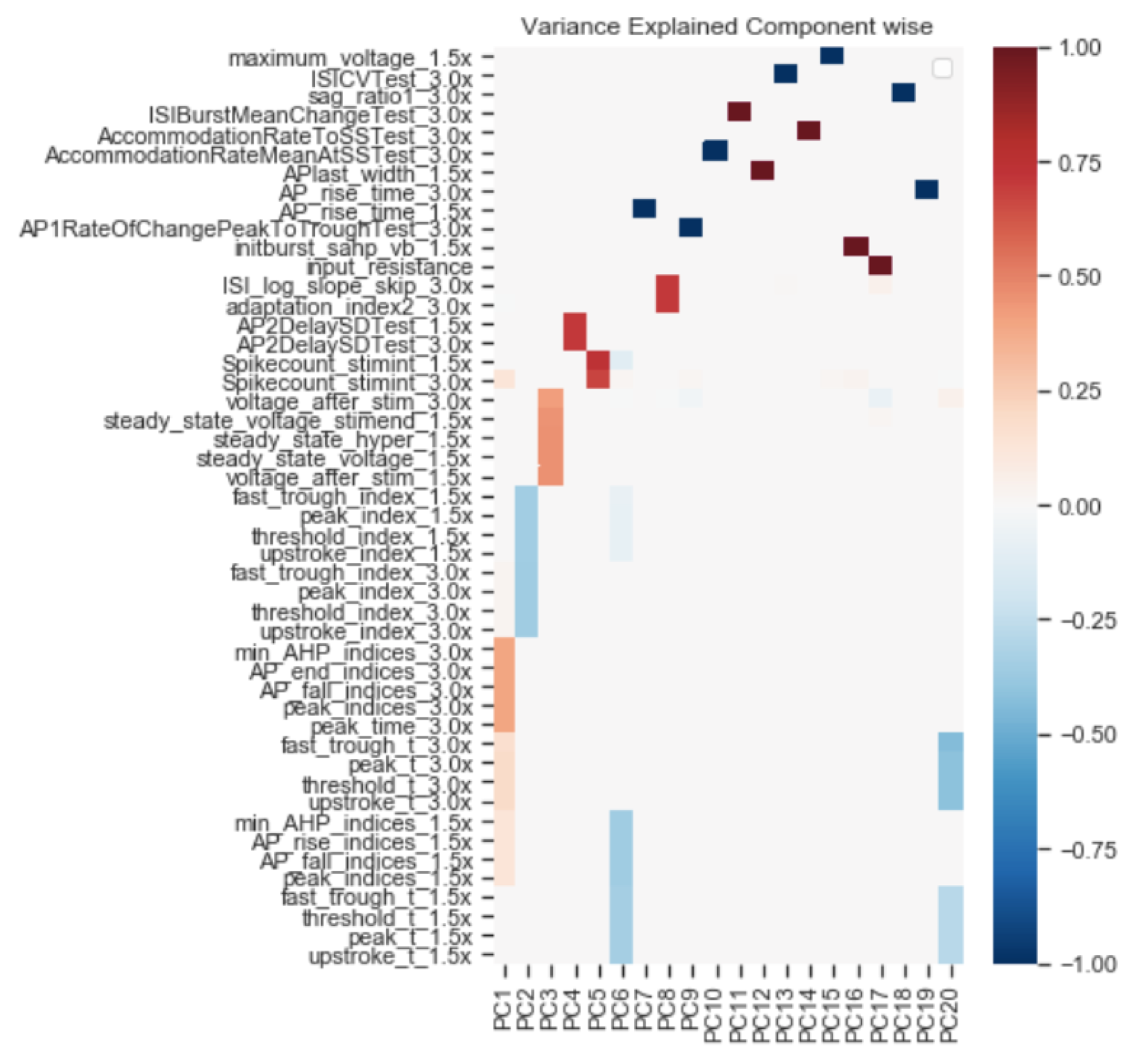
Next Steps
Optimize biophysically accurate models on the features that most disagree between models and experiments.
Interactive Plots:
Can be included or externally linked. dimension_reduced_TSNE
Building the Poster
This poster was created with rmarkdown utilizing a python kernel
The code for this poster lives here To compile this poster do:
BASH terminal Run:
R -e "install.packages('posterdown')"
R -e "rmarkdown::render('template_two.Rmd', \
output_file=\
'CNS_poster_2020.html')"To use a python kernel in this poster:
r, include=FALSE,echo=FALSE}
library(reticulate)
use_python("/anaconda3/bin/python") References:
bibliography: Russell_Prospectus_.bib